A new publication from researchers at the Faculty of Archaeology, Ain Shams University, Cairo, including Dr. Abdelrazek Elnaggar; Proteomics and Metabolomics Research Program, Basic Research Department, Children’s Cancer Hospital, Cairo, Egypt; Department of Physiology, Faculty of Veterinary Medicine, Suez Canal University, Ismailia, Egypt; and The Scientific Research Department, the Metropolitan Museum of Art, New York, USA, is now available and freely accessible in Heritage Science : “Paleoproteomic profiling for identification of animal skin species in ancient Egyptian archaeological leather using liquid chromatography coupled with tandem mass spectrometry (LC-MS/MS)”.
Leatherworking is one of the signifcant industries in ancient Egypt where diferent animal skins (sheep, goat, cow, cattle, and gazelle) were tanned by creating chemical bonding between amino acids of the dermal network of the collagen protein and the vegetal or mineral molecules of tanning material. Te protein information of the archaeological leather could provide signifcant insights into the leather technology, authentication, degradation, preservation needs, and environmental impact over the years. This study aimed at providing guidance for choosing proteomics workfows to analyze leather samples and their capacity to distinguish between unknown archeological species.
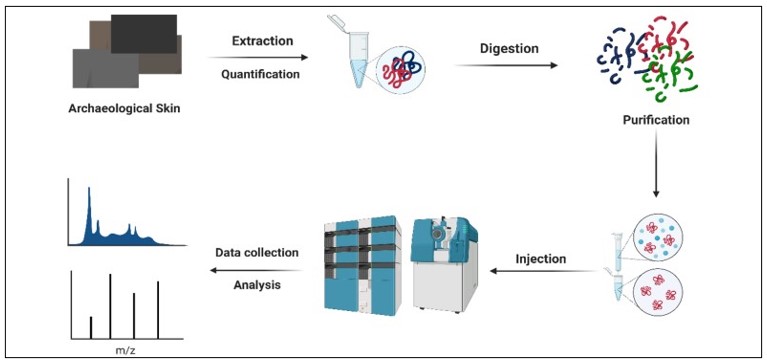
Paleoproteomics has the potential to give a better understanding of the modes of fabrication of ancient materials, their composition, and pathways of degradation, as well as the development of animal fibers through domestication and breeding. Here, we performed shotgun proteomics of archeological animal skin for the first time. The raw output data were analyzed using three diferent software (Proteome Discoverer, Protein Pilot, and Peptide Shaker) with their impeded algorithms.
The study found that the best species identifcation percentage was obtained using protein piolet with protein database. Particularly prevalent and relatively high collagen expression suggests its resistance to degradation, despite the samples’ exposure to environmental and chemical alterations. The success of this case study indicates that further analyses could assist in reworking historical baseline data for putative identifcation of unknown archeological samples
Find more details in the open-access publication here.
